This yeast two-hybrid library was constructed from mouse cDNA that had been previously normalized to preferentially remove abundant cDNAs derived from high-copy-number mRNAs. The normalization process incorporates a Duplex-Specific Nuclease (DSN) treatment and SMART technology, and increases the representation of low-copy-number transcripts in the library. This reduces the number of clones that must be screened to identify positive interactions, and facilitates the identification and characterization of novel protein-protein interactions.
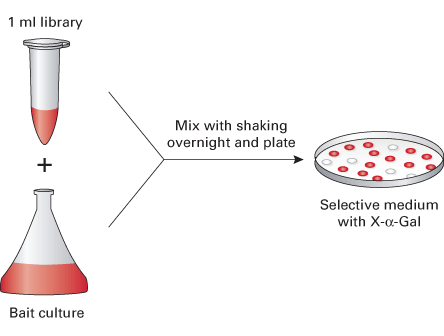
A universal mouse cDNA library transformed into yeast strain Y187. The library can be readily mated to a MATa GAL4 reporter strain, such as AH109 or Y2HGold.
Overview
- By far the easiest way to screen a library for protein-protein interactions
- Screen fewer colonies, detect more interactions
- Universal libraries—broadest gene representation
- No more searching for needles in a haystack