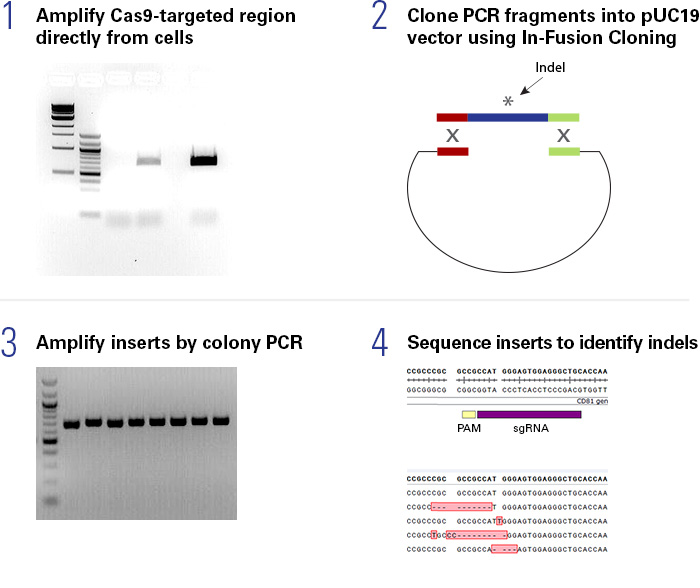
The Guide-it Indel Identification Kit is used for characterization of insertions and deletions (indels) generated by gene editing tools, such as CRISPR/Cas9. This kit contains all of the components needed to amplify, clone, and prepare modified target sites for DNA sequence analysis. This kit uses Terra PCR Direct to amplify targets directly from crude genomic DNA extracts. The resulting pool of fragments, which may contain a variety of indels, are cloned into a prelinearized pUC19 vector using the In-Fusion cloning system. Colony PCR of individual clones using Terra PCR Direct followed by DNA sequencing allows indel characterization.
Overview
- Streamlined method for characterizing the variety of indels introduced by gene editing technologies
- Ultrafast protocol includes Terra PCR Direct Polymerase for PCR amplification directly from cells and the In-Fusion Cloning system for ligation-free cloning in 15 minutes
- Ideal for identifying indels caused by Cas9/CRISPR, TALENs, or zinc-finger nuclease-based gene editing in a cell population
- Complete kit contains all of the components needed to amplify, clone, and prepare modified target sites for DNA sequence analysis
Applications
- Detecting insertions and deletions introduced in mammalian genomic DNA
- Characterizing the variety of indels that are present in a cell population after nuclease-based gene editing