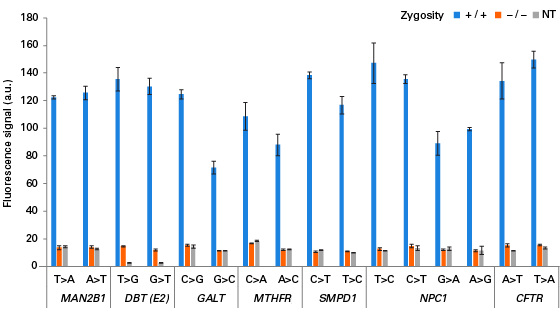
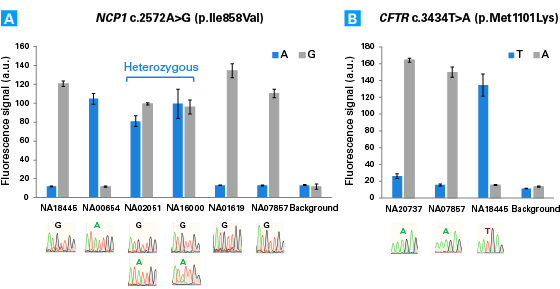
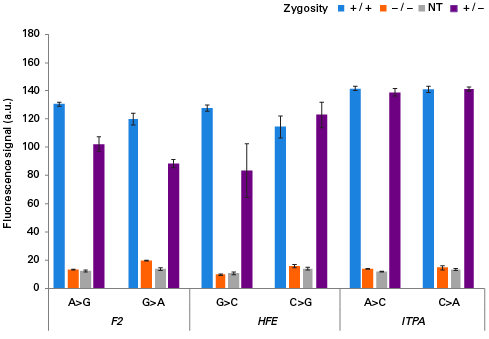
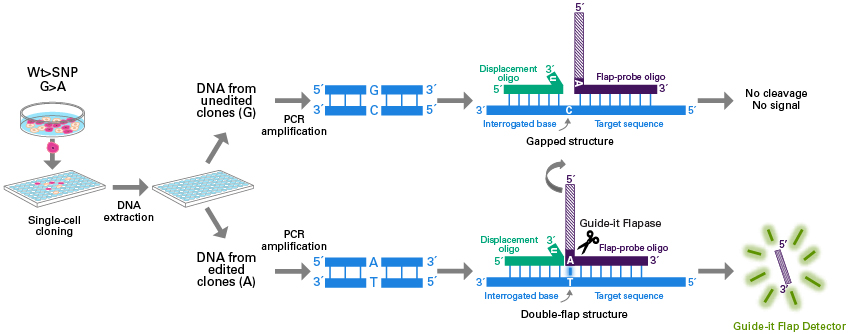
The Guide-it SNP Screening Kit is designed for rapid, high-throughput detection of single-nucleotide substitutions in cell populations that have been edited using technologies such as the CRISPR/Cas9 system. One of the most powerful applications of genome editing technology involves the introduction of single-nucleotide substitutions at specific genomic sites to mimic single-nucleotide polymorphisms (SNPs) implicated in human diseases or to introduce stop codons that yield precise gene knockouts. However, screening hundreds of clones for a single-nucleotide edit remains a challenge, especially in the absence of a phenotype. The Guide-it SNP Screening Kit provides the ability to quickly identify edited clones in 96-well plates, employing a simple and rapid workflow that comprises PCR amplification of the genomic target site followed by an enzymatic assay with a fluorescent readout. The overall workflow takes approximately four hours to complete and uses a standard fluorescence plate reader. The stringency of the assay is such that detection of a fluorescent signal is positively correlated with successful introduction of the desired substitution. This method requires no optimization, and performs comparably regardless of the specific nucleotide substitution, the resulting zygosity of the clone, or the sequence of the targeted locus.
Overview
- High-throughput SNP screening without the need for sequencing
- A simple and rapid workflow that takes just four hours to complete
- Can be used to detect any nucleotide substitution at any genomic locus
-
Identifies edited clones regardless of their zygosity
Applications
- Rapid detection of single-nucleotide substitutions in edited cell populations
- SNP genotyping